Ensembl Release 49!
Yesterday, Ensembl released a new version of the browser and database (version 49). Along with new species, homologue predictions, and new code in our API, there have been changes in how the multiple alignments are done on the whole-genome scale. Have a look at the news for more details.
We are looking forward to release 50! as we are working on some new features. Keep your eye out in August for this next release. A reminder, we will not release another version between now and August, and updates may appear in the Pre! site but not in the main site, for that time.
Please explore features on release 49 such as BLAST which is now configured to align queries against top-level sequences (i.e. chromosomes and scaffolds), and BLAT, a fast alignment program which is now the default selection.
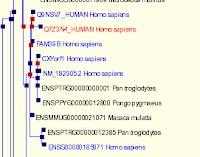
Paralogues are shown in blue in GeneTreeView to help aid your eye.
Upcoming workshops -April
(March workshops are listed in a previous post)
Browser workshops at the VIB Ghent and Leuven (31 Mar - 2 Apr)
Browser workshop (focus: rat) at the ULB Brussels (EURATools) (16 Apr)
Browser workshop at the BCB UCL/Birkbeck (21 Apr)
Module in the EBI roadshow in Poitiers (23, 24 Apr)
API workshop at the Dept. of Genetics, Cambridge (28, 29, 30 Apr)